|
11/6/2022 0 Comments Stage plot pro text on bottom Several experimental advantages have facilitated the exhaustive reconstruction of developmental gene regulatory networks (GRNs) in sea urchin embryos. In particular, we are interested in the predictive value of enhancer transcription estimated by the analysis of the transcription run-on assay PRO-seq, which detects the differential location of paused and elongating RNA Pol II associated with distinct transcriptional regulatory states. Enhancer transcription may facilitate enhancer activity prediction because it represents the end product, possibly subsequent in most cases to chromatin accessibility set in part by H3K27 acetylation, and because it correlates between enhancers and their target promoters. However, the systematic evaluation of the predictive power and redundancy of these genomic marks remains limited. Histone marks such as H3K27ac, chromatin accessibility or transcription initiation facilitate the identification of active enhancers. Various high-throughput reporter assay with particular advantages and limitations allow genome-wide testing of enhancer activity, and, despite recent progress, most TREs remain largely uncharted. Despite their relevance, only a few cis-regulatory modules (CRMs) that constitute the essential transcriptional nodes of developmental gene regulatory networks are functionally understood. In contrast, the oftentimes complex expression of developmental regulatory genes (that is, transcription and signaling factors) is primarily controlled by distal regulatory elements. The expression of inducible genes in unicellular organisms is oftentimes driven by regulatory sequences proximal to the core promoter. Nevertheless, distinct sequences and chromatin features associate with the prevalence of enhancer and promoter activities among TREs. However, this exclusive functional distinction has been blurred by recent evidence that reveals local transcription initiation at enhancers and promoters that modulate the transcription of some other promoters. Traditionally, enhancers have been considered the modulators of distal transcription at core promoters (promoters thereafter), which integrate inputs from enhancers and ‘proximal promoters’ to initiate local transcription. Whereas promoters are TREs easily found by association with the transcription start sites (TSSs) of genes, the identification and functional characterization of TREs distal to TSSs (enhancers and silencers) remains challenging. 
Stage plot pro text on bottom drivers#Transcriptional regulatory elements (TREs) are the primary drivers of differential gene expression during metazoan development. This suggests that the sequence of regulatory mechanisms leading to transcriptional activation have distinct relevance at different levels of the developmental gene regulatory hierarchy deployed during embryogenesis. Furthermore, ATAC- and PRO-seq predictive value is stage dependent for the promoter-overlapping subset. 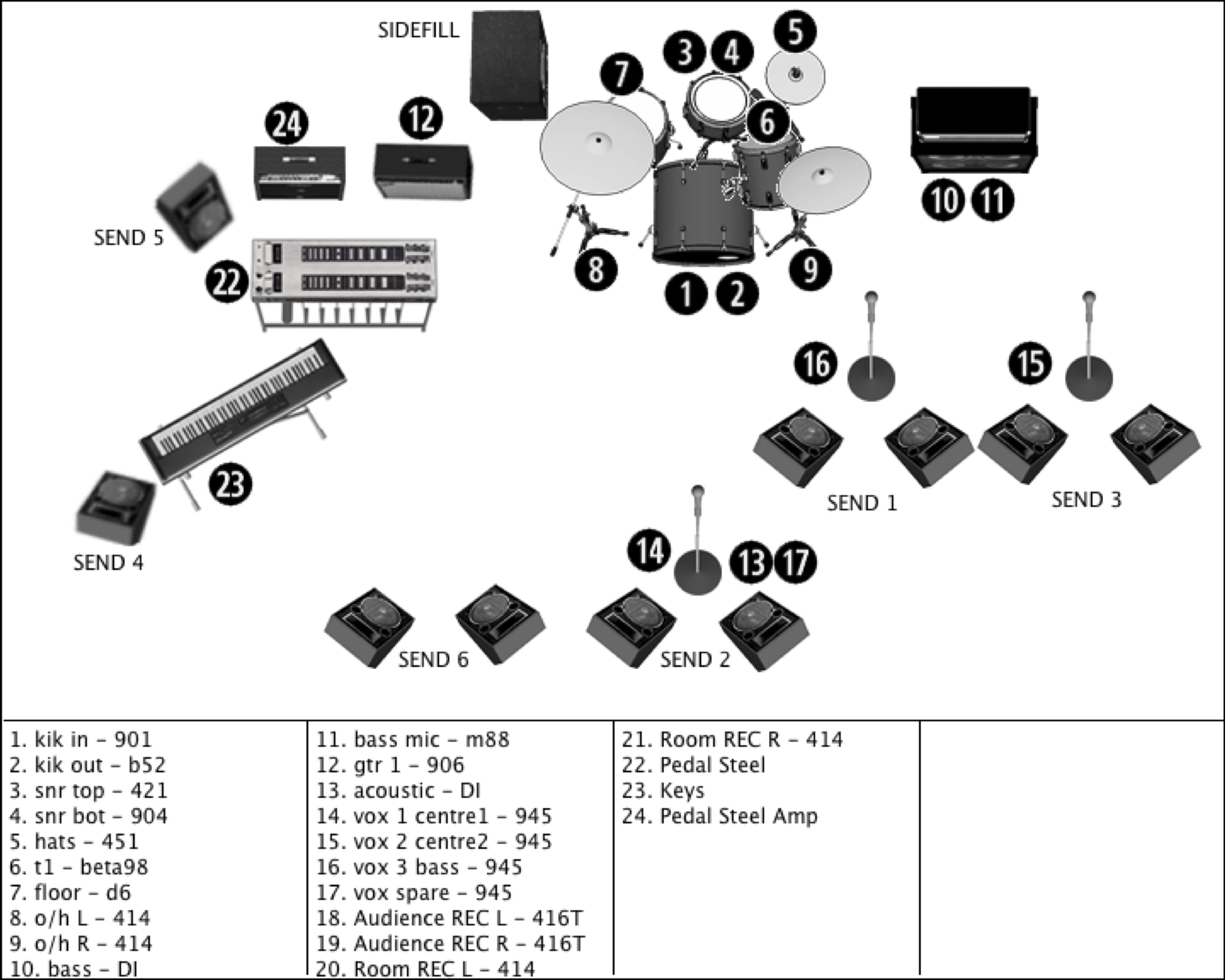
For promoter-overlapping cis-regulatory elements in particular, the distribution of Pol II is the best predictor of enhancer activity in blastula embryos. 
Overall, chromatin accessibility and transcription have substantial power for predicting enhancer activity. Using the three functional genomic data types, machine learning models are trained and tested to classify and quantitatively predict the enhancer activity of several hundred genomic regions previously validated with reporter constructs in vivo. ATAC-seq, PRO-seq, and Pol II ChIP-seq from early and late blastula embryos are manually contrasted with experimental cis-regulatory analyses available in sea urchin embryos, with particular attention to common developmental regulatory elements known to have enhancer and silencer functions differentially deployed among embryonic territories. We evaluate the use of chromatin accessibility, transcription and RNA Polymerase II for their ability to predict enhancer activity of genomic regions in sea urchin embryos. The identification and functional characterization of distal regulatory elements remains challenging, even in tractable model organisms like sea urchins. The transcription of developmental regulatory genes is often controlled by multiple cis-regulatory elements. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
